This is an ongoing project under UCSD's Advanced Robotics and Control Lab (ARCLab), in collaboration with a medical robotics company and physicians at UCSD Health. The goal is to build an autonomous system that can pilot a monocular robotic endoscope through the kidney's collecting system to locate and destroy kidney stones—a procedure known as flexible ureteroscopy.
During ureteroscopy, a flexible scope is passed through the urinary tract into the kidney. Practitioners navigate the scope through the kidney's branching internal structure (the calyces) to find and treat stones. In practice, it is possible for even experienced operators to miss calyces and therefore miss stones, requiring patients to return for follow-up procedures. An autonomous system that can thoroughly and systematically explore every branch would address this directly.
Prior information from CT
Fortunately, a CT scan is typically ordered before the procedure. This gives us a 3D model of the patient's kidney ahead of time, from which we can extract:
- The tree/graph structure of the collecting system — nodes (calyces) connected by edges (infundibula)
- A centerline through each branch for path planning
- The number of junctions at each node (i.e., how many branches split off)
This prior knowledge is the foundation of the navigation approach.
Why SLAM failed
The first attempt at solving this problem was to use foundational SLAM algorithms — specifically DROID-SLAM and VGGT-SLAM — to close the loop between the robot following the precomputed centerline and the SLAM system estimating the robot's true position.
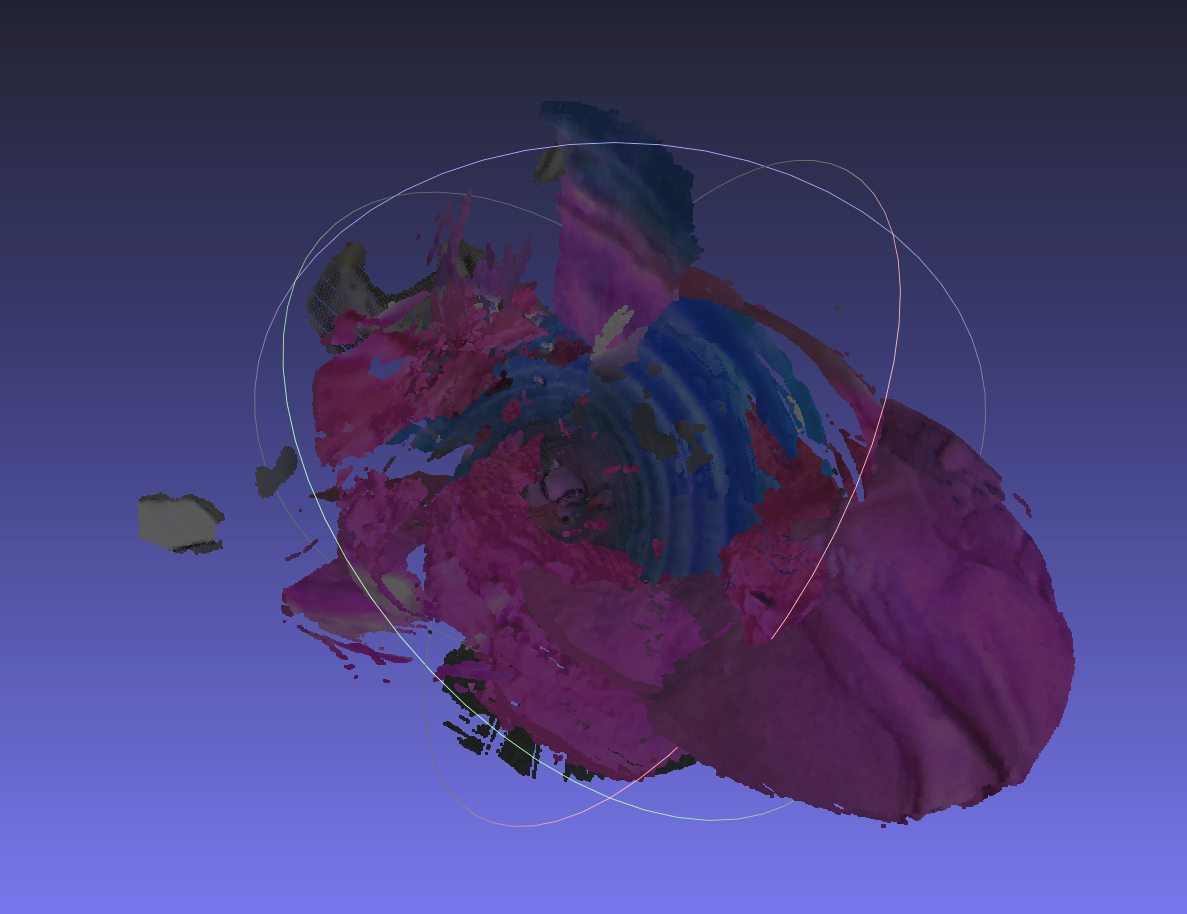

However, the interior of the kidney is featureless, wet tissue. Even with dense reconstruction, these visual SLAM approaches cannot reliably track features in this environment. The lack of texture, combined with specular reflections from the wet mucosal surface, makes feature matching and loop closure extremely fragile.
Key insight: graph-based navigation
One key observation is that we don't necessarily need the precise metric position of the camera. Kidneys — and other complex organs like the lungs — can be modeled as tree structures. We know the graph topology ahead of time from the CT scan: the nodes, the edges, and crucially, the number of branches at each junction.
This means the navigation problem can be reframed:
- If we enter the kidney and reach a junction that we know (from CT) has 3 branches, but the camera currently sees only 2, the system can reason that there must be a 3rd branch it isn't seeing and needs to rotate the camera to find it.
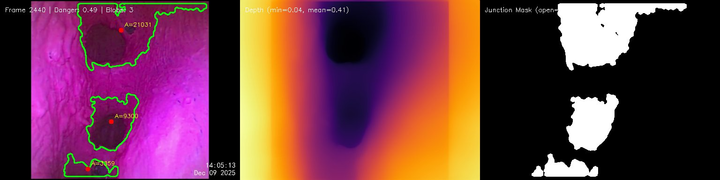
- Rather than dense 3D reconstruction, we only need depth information — enough to determine how many junctions the camera is seeing and to ensure the scope doesn't collide with the tissue walls (not catastrophic if it does, but should be avoided).
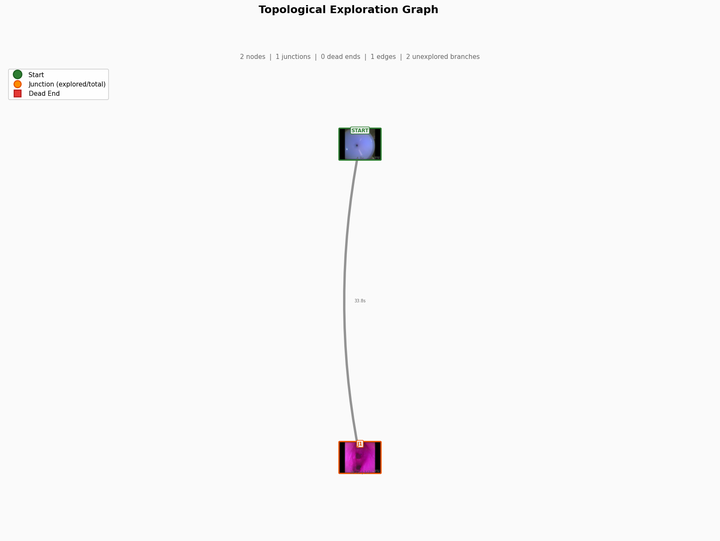
This reduces the problem from full SLAM to depth-based junction detection combined with graph-level localization: answering "which node in the known graph are we at?" rather than "what is our exact 6-DoF pose?"
Current research
Research is currently focused on two threads:
-
Minimal sensing for agentic navigation — determining what tools and information are sufficient for an agentic system to navigate the kidney's graph structure. Key questions include how to localize within the node graph using depth and junction count, and how to distinguish revisited locations from new ones to ensure complete coverage.
-
Procedural ureteroscopy simulator (endonav-sim) — a seeded simulator that generates a new anatomically-grounded kidney on every run, so navigation controllers are evaluated against diverse anatomies rather than one canonical phantom.
Every seed produces a unique Sampaio Type A1 pelvicalyceal tree with sampled minor-calyx count (7–13), lower-pole infundibula count (3–7), infundibulopelvic angle, and ureter narrowings. Kidney stones are placed with epidemiologically-accurate composition (~80% calcium oxalate), lognormal sizing (2–20 mm), and gravity-biased lower-pole distribution, and both clinical treatment modes are modeled — basket capture for stones ≤3.5 mm and laser fragmentation for stones ≤15 mm.
The ureteroscope itself is simulated with honest dynamics: encoder quantization, deflection dead-zones, tendon-sheath backlash hysteresis, shaft buckling, retraction slip, and dead-reckoned pose drift — the exact failure modes a real controller has to contend with. Rendering uses an EndoPBR-style coaxial light model (GGX/Cook–Torrance specular, warm subsurface diffuse, ACES tonemap) through phantom-camera optics with SSAA and chromatic aberration, running at 260+ fps on a single kidney with stones.
Repositories
- Procedural ureteroscopy simulator: https://github.com/TylerFlar/endonav-sim
- Depth estimation and branch detection experiments: https://github.com/TylerFlar/endonav-exploration